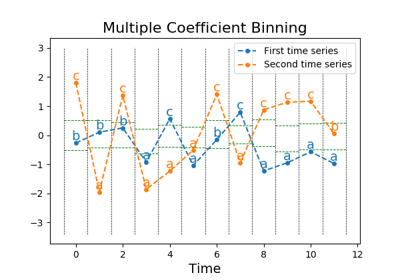
pyts.approximation.MultipleCoefficientBinning¶
-
class
pyts.approximation.MultipleCoefficientBinning(n_bins=4, strategy='quantile', alphabet=None)[source]¶ Bin continuous data into intervals column-wise.
Parameters: - n_bins : int (default = 4)
The number of bins to produce. It must be between 2 and
min(n_samples, 26).- strategy : str (default = ‘quantile’)
Strategy used to define the widths of the bins:
- ‘uniform’: All bins in each sample have identical widths
- ‘quantile’: All bins in each sample have the same number of points
- ‘normal’: Bin edges are quantiles from a standard normal distribution
- ‘entropy’: Bin edges are computed using information gain
- alphabet : None, ‘ordinal’ or array-like, shape = (n_bins,)
Alphabet to use. If None, the first n_bins letters of the Latin alphabet are used if n_bins is lower than 27, otherwise the alphabet will be defined to [chr(i) for i in range(n_bins)]. If ‘ordinal’, integers are used.
References
[Rfea62cc40411-1] P. Schäfer, and M. Högqvist, “SFA: A Symbolic Fourier Approximation and Index for Similarity Search in High Dimensional Datasets”, International Conference on Extending Database Technology, 15, 516-527 (2012). Examples
>>> from pyts.approximation import MultipleCoefficientBinning >>> X = [[0, 4], ... [2, 7], ... [1, 6], ... [3, 5]] >>> transformer = MultipleCoefficientBinning(n_bins=2) >>> print(transformer.fit_transform(X)) [['a' 'a'] ['b' 'b'] ['a' 'b'] ['b' 'a']]
Attributes: - bin_edges_ : array, shape = (n_bins - 1,) or (n_timestamps, n_bins - 1)
Bin edges with shape = (n_bins - 1,) if
strategy='normal'or (n_timestamps, n_bins - 1) otherwise.
Methods
__init__(self[, n_bins, strategy, alphabet])Initialize self. fit(self, X[, y])Compute the bin edges for each feature. fit_transform(self, X[, y])Fit to data, then transform it. get_params(self[, deep])Get parameters for this estimator. set_params(self, \*\*params)Set the parameters of this estimator. transform(self, X)Bin the data. -
__init__(self, n_bins=4, strategy='quantile', alphabet=None)[source]¶ Initialize self. See help(type(self)) for accurate signature.
-
fit(self, X, y=None)[source]¶ Compute the bin edges for each feature.
Parameters: - X : array-like, shape = (n_samples, n_timestamps)
Data to transform.
- y : None or array-like, shape = (n_samples,)
Class labels for each sample. Only used if
strategy='entropy'.
-
fit_transform(self, X, y=None, **fit_params)¶ Fit to data, then transform it.
Fits transformer to X and y with optional parameters fit_params and returns a transformed version of X.
Parameters: - X : numpy array of shape [n_samples, n_features]
Training set.
- y : numpy array of shape [n_samples]
Target values.
- **fit_params : dict
Additional fit parameters.
Returns: - X_new : numpy array of shape [n_samples, n_features_new]
Transformed array.
-
get_params(self, deep=True)¶ Get parameters for this estimator.
Parameters: - deep : bool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
Returns: - params : mapping of string to any
Parameter names mapped to their values.
-
set_params(self, **params)¶ Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as pipelines). The latter have parameters of the form
<component>__<parameter>so that it’s possible to update each component of a nested object.Parameters: - **params : dict
Estimator parameters.
Returns: - self : object
Estimator instance.